Bayesian Phylogeography
What is Phylogeny?
A simple example of a phylogeny is a family tree where the leaves of the tree represent the children in the family and branches represent relationships between parents and children. The tree represents different clans inside a larger family.
Likewise, a language phylogeny is a tree representation of the closeness of various languages. Dutch and Flemish are sister-languages that have a very close common ancestor. English is a cousin language, which is a bit further away from Dutch and Flemish. The Scandinavian languages are far cousins.
The place where lines come together in the language ‘family tree’ represent older languages that gave rise to the child languages in the tree.
Below, an example of a phylogeny for ancient languages (From Ringe).
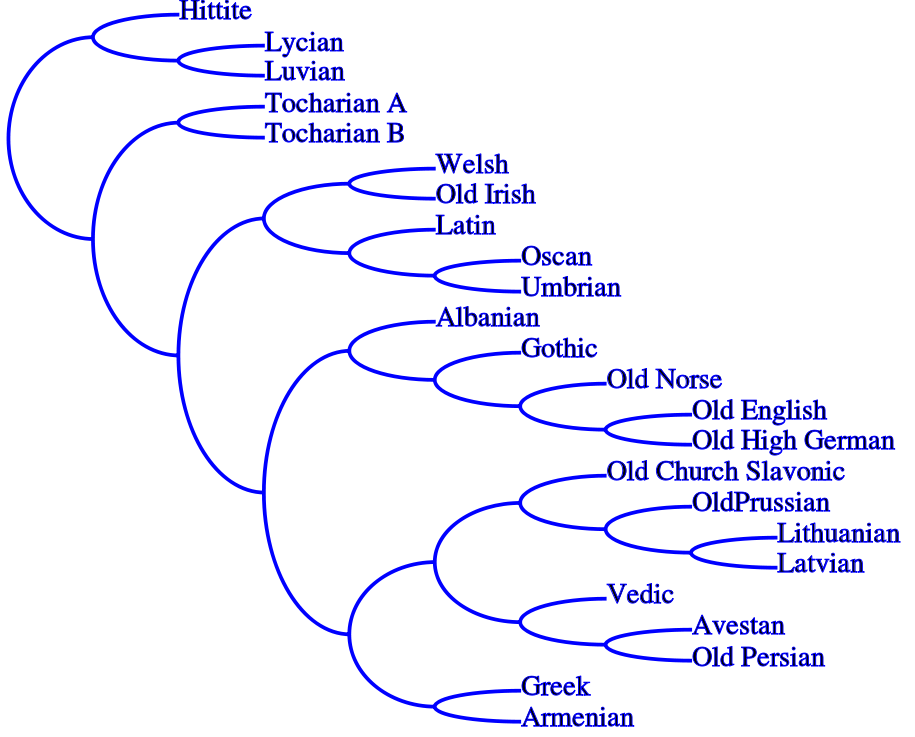
How do we infer phylogenies/family trees?
Family trees can be inferred by looking at the DNA of the children in the family. Siblings have much more of their DNA in common than far cousins. Siblings have by definition parents in common. However, we do not have the DNA of parents or any other predecessor available. Determining siblings and their parents is typically quite straightforward, but the further we go in the past, the more uncertain we are about the relationships.
It is a bit like looking at a tree where we can see the leaves covering the branches of the tree and we want to reconstruct the branches and trunk of the tree. Typically, it is easy to reconstruct the twigs ending in the leaves, but further down the tree uncertainty about the structure of the tree increases.
What is Bayesian Inference?
Bayesian inference is a method based on the idea that before we analyse any data we already have some view of the world. This so called ‘prior’ belief consists of information such as the date that Latin was spoken. Another example is historical sources indicating differentiation of Lithuanian and Latvian in the 7th century as Proto-Latvian and Proto-Lithuanian tribes emerged. This means that Latvian and Lithuanian must be siblings in the tree. Furthermore we have an indication of the time for their parents.
After adding data and a model, we reach a ‘posterior’ belief of the world. In our analysis, we use language and location data and a phylogeographical model as explained further below.
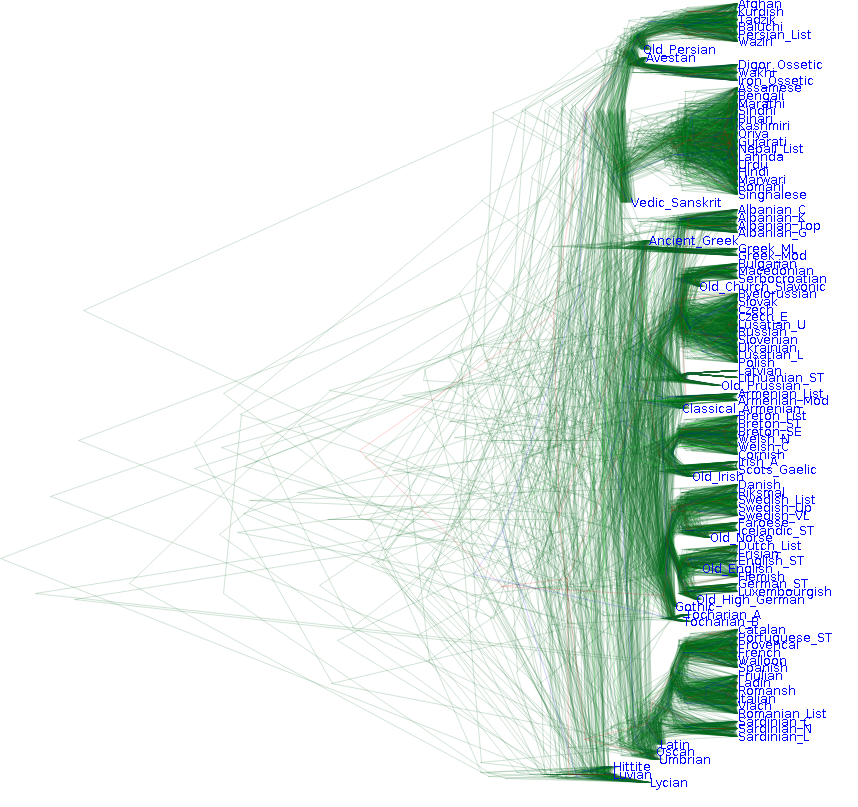
To see how this works, the following image shows our prior belief over phylogenies over the languages. Note that it shows a cloud of trees which represent a distribution of possible trees. Bayesian inference does not result in a single viewpoint of the world, but rather a distribution with some scenarios being more likely than others.
Also note that there is some structure grouping some languages together reflecting our prior belief. However, higher up in the tree there is large uncertainty around the structure and timing of the tree.
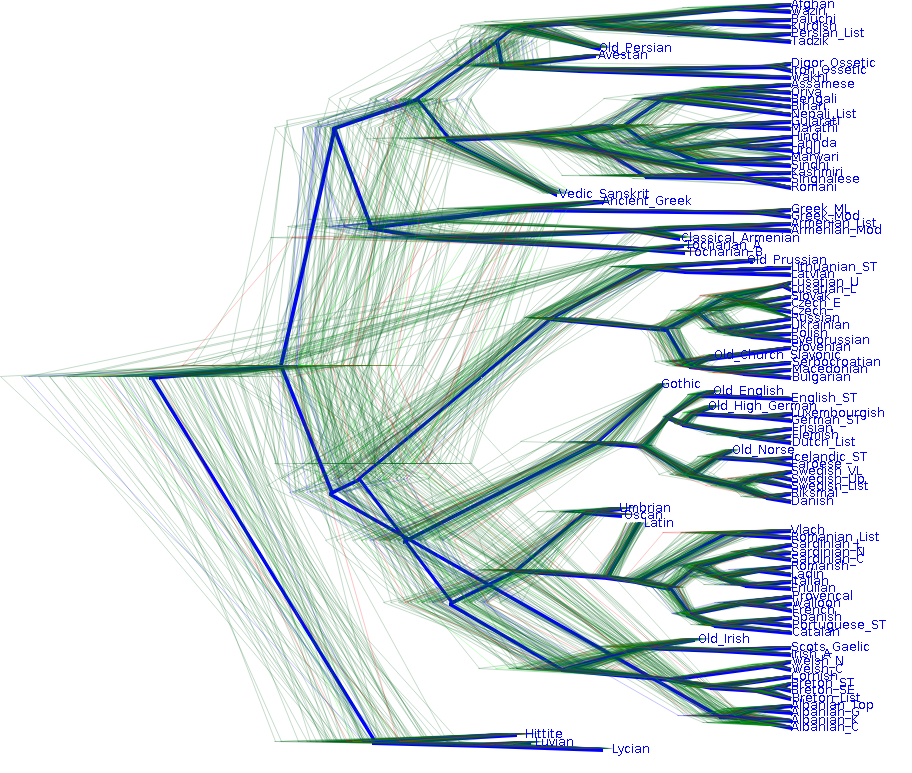
After analysing our language and location data, we obtain a posterior distribution of trees that is visualised below. The blue line represent a summary tree which is a good reflection of our belief taking the prior and data in account. Clearly, the data considerably reduced uncertainty about the structure of the tree. However, note that there is still some uncertainty left both in the timing of the tree and in the structure as well.
What is Phylogeography?
Phylogeography is a method that was initially developed to examine the geographic spread of one particular part of the tree of life – viruses. It combines geographic data with sequence data, DNA sequence in the case of viruses and cognates in this study. It can be used to answer questions like where did the last H5N1 strain come from?
These methods don’t just infer family-tree-like relationships between the organisms in question, but also trace back along the lineages of the tree to infer the geographic location at the root of the tree – the geographic origin point.
What is Bayesian Phylogeography?
Bayesian Phylogeography combines the Bayesian method of inference (see above) with phylogeography. Because the Bayesian method allows for easy incorporation of different kinds of prior information, we can specify calibration dates for various internal nodes in the tree as well as regions of origin of languages.
The result of a Bayesian phylogeographical analysis gives us a distribution for the locations of the internal nodes in the tree as well as a distribution of the location of the origin of the tree.
What are the differences between geographic models?
We considered different geographical models that are based on random ‘drunken man’ walks on a map. There is a large body of theoretical work on random walks and the walks are restricted by their end points; the locations where the languages are spoken.
The main difference between the two models we considered is that one is based on continuous space and the other discrete space, that is, the first model allows steps of arbitrary size and direction, while the second model allows steps only on a fine grid. At first sight, the second model sounds restrictive, but it means we can distinguish between land and water, something that is not quite so straightforward with the first model.
In fact, we can model how keen or reluctant migrants are to enter water, and then travel further on over sea. To see how land/water influences the rate of migration, below some animations of the landscape aware model for different areas. The color intensity changes when population density changes with light blue for high densities and dark blue for low densities. First, Scotland with a lot of water around it. The animation puts the population initially in Scotland from where migration starts out to England and over seas towards the rest of Great Britain and mainland Europe. Clearly, migration over sea is a lot faster, but the amount of migration is a lot lower.
The animation for migration from a block in Uzbekistan shows what happens when there is almost no water to help or hinder migration. Finally, migration from a block in Turkey shows how the Mediterranean sea can quickly be reached, while migration over land is a lot slower.
Animation starting in Scotland
Animation starting in Uzbekistan
Animation starting in Turkey

